|
Vector Laboratories
biotinylated igg secondary antibodies Biotinylated Igg Secondary Antibodies, supplied by Vector Laboratories, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/biotinylated igg secondary antibodies/product/Vector Laboratories Average 94 stars, based on 1 article reviews
biotinylated igg secondary antibodies - by Bioz Stars,
2026-03
94/100 stars
|
Buy from Supplier |
|
Sino Biological
c fasciculata C Fasciculata, supplied by Sino Biological, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/c fasciculata/product/Sino Biological Average 90 stars, based on 1 article reviews
c fasciculata - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
BIO-CAT Inc
pcdh-cmv-mcs-ef1-hygro Pcdh Cmv Mcs Ef1 Hygro, supplied by BIO-CAT Inc, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pcdh-cmv-mcs-ef1-hygro/product/BIO-CAT Inc Average 90 stars, based on 1 article reviews
pcdh-cmv-mcs-ef1-hygro - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
Amersham Pharmacia Biotech Ltd
psk-cat vector controls Psk Cat Vector Controls, supplied by Amersham Pharmacia Biotech Ltd, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/psk-cat vector controls/product/Amersham Pharmacia Biotech Ltd Average 90 stars, based on 1 article reviews
psk-cat vector controls - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
Promega
59 and 39 src1a promoter deletion cat expression vectors 59 And 39 Src1a Promoter Deletion Cat Expression Vectors, supplied by Promega, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/59 and 39 src1a promoter deletion cat expression vectors/product/Promega Average 90 stars, based on 1 article reviews
59 and 39 src1a promoter deletion cat expression vectors - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
VectorBuilder GmbH
a grna expression vector targeting human mmp3 (cat. no. vb170623-1031qnn)  A Grna Expression Vector Targeting Human Mmp3 (Cat. No. Vb170623 1031qnn), supplied by VectorBuilder GmbH, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/a grna expression vector targeting human mmp3 (cat. no. vb170623-1031qnn)/product/VectorBuilder GmbH Average 90 stars, based on 1 article reviews
a grna expression vector targeting human mmp3 (cat. no. vb170623-1031qnn) - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
Promega
stk3 nanobret kinase assays  Stk3 Nanobret Kinase Assays, supplied by Promega, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/stk3 nanobret kinase assays/product/Promega Average 90 stars, based on 1 article reviews
stk3 nanobret kinase assays - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
Shanghai GenePharma
pcdna empty vector (cat. no. g01001, concentration: 10 8 tu/ml)  Pcdna Empty Vector (Cat. No. G01001, Concentration: 10 8 Tu/Ml), supplied by Shanghai GenePharma, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pcdna empty vector (cat. no. g01001, concentration: 10 8 tu/ml)/product/Shanghai GenePharma Average 90 stars, based on 1 article reviews
pcdna empty vector (cat. no. g01001, concentration: 10 8 tu/ml) - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
BIO-CAT Inc
pcdna3 expression vector  Pcdna3 Expression Vector, supplied by BIO-CAT Inc, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pcdna3 expression vector/product/BIO-CAT Inc Average 90 stars, based on 1 article reviews
pcdna3 expression vector - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
BIO-CAT Inc
expression vectors mutant variants c4- nonc4-nadp-me  Expression Vectors Mutant Variants C4 Nonc4 Nadp Me, supplied by BIO-CAT Inc, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/expression vectors mutant variants c4- nonc4-nadp-me/product/BIO-CAT Inc Average 90 stars, based on 1 article reviews
expression vectors mutant variants c4- nonc4-nadp-me - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
Promega
cat expression vectors pcatenhancer  Cat Expression Vectors Pcatenhancer, supplied by Promega, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/cat expression vectors pcatenhancer/product/Promega Average 90 stars, based on 1 article reviews
cat expression vectors pcatenhancer - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
|
Montana Molecular
baculovirus vector pseudotyped with spike m2 green reporter (cat # c1122g)  Baculovirus Vector Pseudotyped With Spike M2 Green Reporter (Cat # C1122g), supplied by Montana Molecular, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/baculovirus vector pseudotyped with spike m2 green reporter (cat # c1122g)/product/Montana Molecular Average 90 stars, based on 1 article reviews
baculovirus vector pseudotyped with spike m2 green reporter (cat # c1122g) - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) uMAP projection of cell density by control and BRCA1 +/mut fibroblasts. b) Volcano plot with all differentially expressed genes between control and BRCA1 +/mut fibroblasts. Top 50 BRCA1 +/mut genes were used to define pre-CAF signature. Top 50 control genes define our control Fibroblast Signature. c) Gene signature scoring of all control and BRCA1 +/mut fibroblasts for cancer associated fibroblast (CAF) and iCAF signatures. P-value determined by welch two sample t-test. d) Overall survival Kaplan-Meier analysis of n=1764 patients with a high (n=970) and low (n=794) expression of the pre-CAF signature. e) Overall survival Kaplan-Meier analysis of n=1764 patients with a high (n=1197) and low (n=567) expression of the control fibroblast signature. f) Boxplot of MMP3 expression in control and BRCA1 +/mut fibroblasts. P values are determined using a Mann-Whitney test. g) In situ immunofluorescence (IF) analysis of lobular and ductal regions of breast epithelium in control and BRCA1 +/mut human tissue sections. Scale bar = 50 µm. h) Dot plot showing percentage of MMP3-positive stromal cells in control (blue) and BRCA1 +/mut (red) samples as quantified from IF stainings. P -values were determined using unpaired t-tests.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Expressing, MANN-WHITNEY, In Situ, Immunofluorescence

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) Pre-CAF gene signature scoring in fibroblasts from individual patients. Libraries with representation of less than 250 fibroblasts were excluded. P -value was determined by a t-test. a) Example immunofluorescence microscopy images from ductal and lobular regions in control tissue samples. Values represent the quantified percentage of stromal cells positive for MMP3. Scale bar = 50 µm. b) Example immunofluorescence microscopy images from ductal and lobular regions in BRCA1 +/mut tissue samples. Values represent the quantified percentage of stromal cells positive for MMP3. Scale bar = 50 µm.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Immunofluorescence, Microscopy

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) Schematic primary human 3D co-culture using FACS-isolated epithelial cells and fibroblasts modulated using lentiviral transduction. b) Representative images depicting green fluorescent protein (GFP) expression in transduced fibroblasts in close proximity with epithelial organoids (arrow). Scale bar = 100 μm. c) Co-cultures (5 days) of 4,000 breast epithelial cells seeded alone (no fibroblasts) or with 1x10 4 control fibroblasts transduced with lentivirus (+GFP) or transduced to express MMP3 and GFP (+MMP3). Western blots show increased expression of MMP3 in cells and cultured supernatant of +MMP3 fibroblasts (left). Representative merged bright field and GFP images of co-cultures (scale bar= 400 μm) with arrows indicating mammospheres (GFP-negative). Bar charts (right) represent number of mammospheres quantified per well; values expressed as mean ± SD from three separate experiments with three triplicate wells from each. P -values were determined using unpaired t-tests. d) Co-cultures (5 days) of 4,000 breast epithelial cells seeded alone (no fibroblast) or with 1x10 4 BRCA1 +/mut fibroblasts transduced with lentivirus to express CRISPR- Cas9 and MMP3 gRNA (-MMP3) or GFP only (+GFP) vectors. Western blots show decreased expression of MMP3 in cells and medium of MMP3 knock out fibroblast cultures (left). Representative overlay bright field and GFP images of co-cultures (scale bar= 400 μm) with arrows indicating mammospheres (GFP-negative). Bar charts (right) represent number of mammospheres quantified in each well; values expressed as mean ± SD from triplicates of three independent experiments. P - values were determined using unpaired t-tests. e) 1x10 4 FACS-isolated epithelial cells from patient sample “Control 36” were seeded in Matrigel and treated with 0.5 µg/mL or 1 μg/mL recombinant MMP3. Number of mammospheres was quantified after 5 and 10 days. Representative bright field images of mammospheres after 10 days of culture (scale bar= 400 μm). Bar chart values are expressed as mean ± SD from triplicates of three independent experiments. P -values were determined using unpaired t-tests. f) Schematic of mouse model for evaluating effects of stromal fibroblasts (Control 27) on BRCA1-mediated breast tumorigenesis in vivo . g) Images of dissected tumors after six weeks of growth with reported tumor formation efficiencies. Scale bar= 1 cm. P -values were determined using a one- sided Fisher’s Exact test. h) Volumes of dissected tumors. Values are represented as mean ± SD. control n= 4; +GFP n=8, MMP3 n=12. P -values were determined using unpaired t-tests. i) Masses of dissected tumors. Values are represented as mean ± SD. control n= 4; +GFP n=8, +MMP3 n=12. P -values were determined using unpaired t-tests.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Co-Culture Assay, Isolation, Transduction, Expressing, Western Blot, Cell Culture, CRISPR, Knock-Out, Recombinant, In Vivo

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) FACS plots showing gating strategy for isolation of transduced human fibroblasts with GFP fluorescence in forward and side scatter, singlets gate, GFP gate. b) Representative images of primary mammary epithelial cells cultured alone (control) or with primary mammary fibroblasts transduced with lentivirus to express GFP only (+GFP) or GFP and MMP3 (+MMP3) in Matrigel for 5 days. Fibroblasts are distinguished from epithelial spheres (GFP negative) with GFP fluorescence. Scale bar = 400 μm. c) Quantification of spheres. Values are represented as mean ± SD calculated from nine separate wells per group. P-values were determined by unpaired t-tests. d) Mean values of sphere counts pooled from 3 separate experiments from (b) and . Values are represented as mean ± SD. Statistical significance between all groups was determined with a one-way ANOVA test. e) 10x10 4 FACS-isolated epithelial cells from 4 additional patient samples were seeded in seeded in Matrigel and treated with 0.5 µg/mL or 1 μg/mL MMP3 and spheres were counted after 5 and 10 days. Representative bright field images of mammospheres after 10 days of culture (scale bar= 400 μm). Bar chart values are represented as mean ± SD from triplicates from three separate experiments. P - values were determined using unpaired t-tests. f) Bar graph depicting the fold change in sphere number compared to control (dotted red line) after 10 days of culture with human recombinant MMP3. Bar graph values are expressed as mean ± SD from 15 independent experiments total (5 different patient samples with 3 separate experiments each). g) Primary human breast fibroblasts FACS-isolated from patient sample “Control 27” were transduced to express mouse MMP3 (mMMP3) or GFP only. qPCR analysis was performed on the transduced fibroblasts in two separate trials with three replicates per group. Amplification plot is shown with the difference in the normalized reporter value of the experimental reaction minus the normalized reporter value generated by the instrument (ΔRn) on the y-axis and the cycle number on the x-axis.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Isolation, Fluorescence, Cell Culture, Transduction, Recombinant, Amplification, Generated

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) Unbiased clustering using uMAP projection of all patient epithelial cells. Cells are labeled by mammary epithelial cell state classification as indicated. b) uMAP feature plots displaying single cell energy in facetted plots for control (upper plot) and BRCA1 +/mut cells (lower plot). c) scEnergy distributions across basal, luminal 1, and luminal2 cell types from control and BRCA1 +/mut samples. Mean scEnergy values from individual patients are plotted. P-value determined by welch two sample t-test. d) Volcano plot displaying genes differentially expressed between control and BRCA1 +/mut luminal 1 epithelial cells. e) Gene signature scoring of luminal 1 cells from control and BRCA1 +/mut epithelial cells for basal, myoepithelial, luminal 2-AREG, and luminal 2-MUCL1 marker gene signatures. P-value determined by Welch two sample t-test. f) In situ immunofluroescence analysis of KRT14/KRT19-double positive cells of lobular and ductal regions in control and BRCA1 +/mut tissues with representative images shown. Bar chart (bottom left) indicates the percentage of K14/K19-double positive cells per field of view in control (n=6) and BRCA1 +/mut (n=6). Values are expressed as mean ± SD quantified from at least 5 random fields per patient sample. P -value was determined using an unpaired t-test. g) Single cell Western blot (scWB)-based quantification of the percentage of KRT23+ in FACS-isolated luminal epithelial cells from n=3 control, and n=3 BRCA1 +/mut . Images are representative regions of scWB chips post electrophoresis and antibody probing. Bar chart values are represented as mean ± SD from at least 1000 cells/individual; n=3 control, and n=3 BRCA1 +/mut . h) Schematic workflow for analyzing co-transplant tumors from in vivo study described in -i for KRT8 and KRT5 expression. i) Representative immunofluorescence images of sections from tumors produced from +GFP control or +MMP3 fibroblast transplants. Yellow staining indicates tumor cells double positive for KRT8 and KRT5. Scale bar= 50 µm. j) Bar chart is a quantification of the percent of KRT5 cells positive from KRT8. Values are represented as mean ± SD from counts from three different tumors per group with at least five random fields per tumor tissue. P -value was determined by an unpaired t-test.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Labeling, Marker, In Situ, Western Blot, Isolation, Electrophoresis, In Vivo, Expressing, Immunofluorescence, Produced, Staining
Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
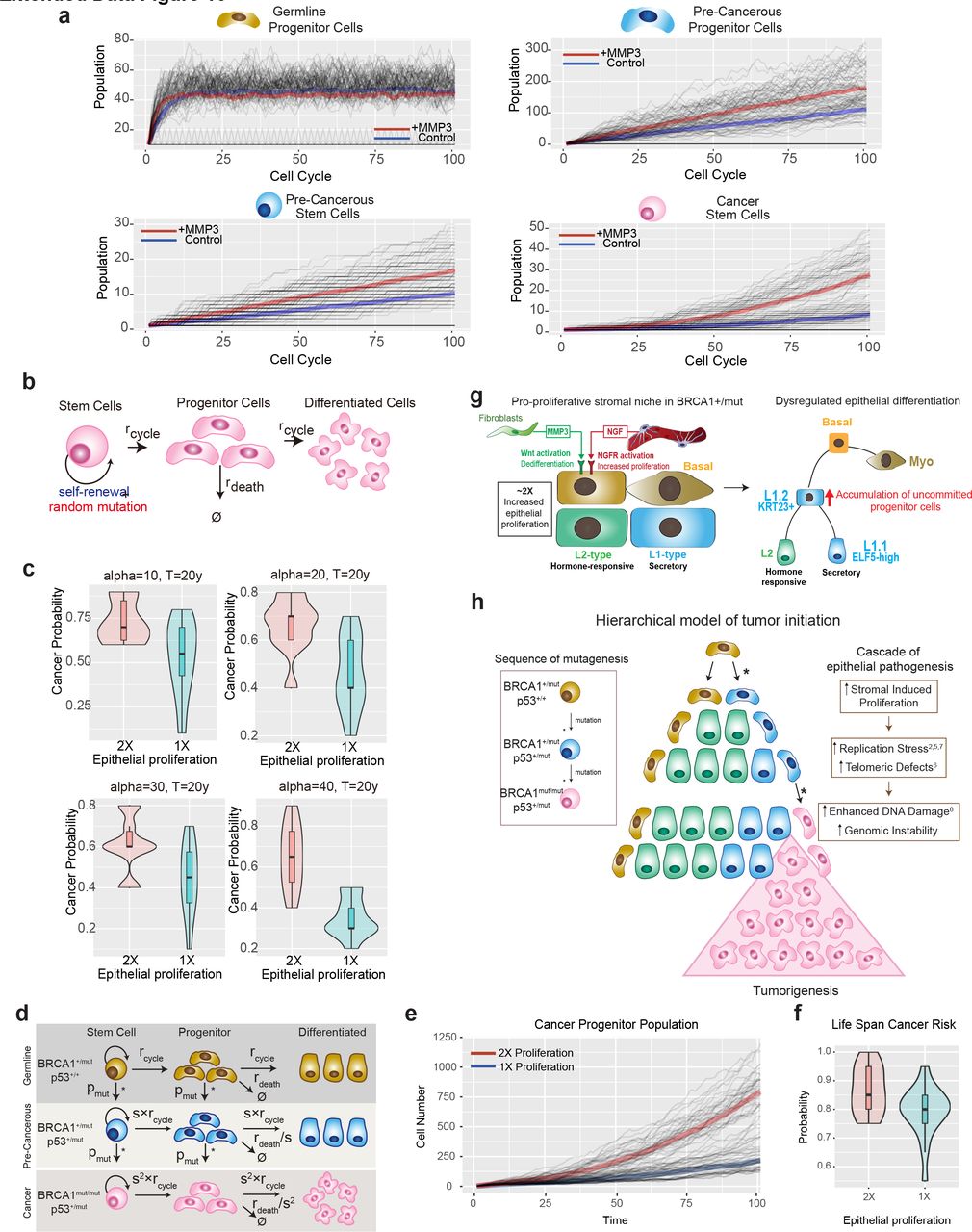
Figure Lengend Snippet: a) The comparison of simulated cell population dynamics in 2 fold increased proliferation and control groups for germline progenitor cells (top left), pre-cancerous progenitor cells (top right), pre-cancerous stem cells (bottom left), and cancer stem cells (bottom right). Thick lines: The averaged population dynamics of 2 fold increased proliferation (red) and control group (blue). Gray thin lines: The stochastic simulation trajectories (sample n=50 for each group). b) Schematic model of the assumptions and parameters used to simulate acquisition of random mutations and consequential fitness change in BRCA1+/mut cells. rcycle is the base line cell division rate, rdeath is the cell date rate. Parameters are further defined in Supplementary Table 10 . c) The robustness of results with respect to parameter alpha in the double-exponential distribution of fitness change ( Supplementary Table 10 ) in the calculation of risk ratio with or without MMP3. The simulation is performed in the time range of 20 years. d) Schematic model illustrating the assumptions and parameters used to simulate the sequential mutations in oncogenes in BRCA1+/mut cells. rcycle is the base line cell division rate, rdeath is the cell date rate, s = proliferation scale factor, pmut is the probability of acquiring a mutation in a driver oncogene. Parameters are further defined in Supplementary Table 11 . e) Comparison between cancer progenitor population dynamics as predicted by a hierarchical model Thick lines: The averaged population dynamics of proliferation in a population with a 2 fold increase in prolifersation and control group (blue). Gray thin lines: The stochastic simulation trajectories (sample n=50 for each group). f) Comparison of the predicted risk ratio of cancer initiation between 2-fold (red) and 1- fold epithelial proliferation rate (blue) in BRCA1 mutation carriers over human life span. The samples are collected from the simulation of N=40 patients in two groups, with the risk ratio of each patient estimated from n=20 simulations of a random mutation model . Violin plots show the distribution of risk ratios over N=20 patients in each group and boxplots indicate median and 25% and 75% quantiles respectively. Wilcoxon Test: *p=0.011. g) Schematic illustrating the concept of a pro-proliferative stromal niche in BRCA1 +/mut breast breast tissues. BRCA1 +/mut stromal cells overepress pro-proliferative cues including NGF in pericytes and pro-tumorigenic MMP3 in fibroblasts (right side). We propose that these act in concert during the pre-neoplastic phase to promote the expansion of a subset of uncommitted luminal progenitor cells as potential cancer cells of origin (left side). h) Concept illustration of the hierarchical model of cancer initiation. Sequence of mutations are indicated in differently colored cells in box on the left, * represent a mutagenic event. Center schematic summarizes outcome of mathematical modeling results indicating expansion of cancer progenitors and ultimately leading to tumorigenesis. Cascade of epithelial cell-intrinsic events promoting tumorigenesis in BRCA1 +/mut is shown on the right. Due to increased stromal cell-induced proliferation and replication stress, BRCA1 +/mut breast epithelial stem cells accumulate mutations and become genomically instable, which increases the likelihood of driver mutations, and ultimately tumor initiation.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Mutagenesis, Sequencing

Journal: Molecular Medicine Reports
Article Title: circSAMD4A participates in the apoptosis and autophagy of dopaminergic neurons via the miR-29c-3p-mediated AMPK/mTOR pathway in Parkinson's disease
doi: 10.3892/mmr.2021.12179
Figure Lengend Snippet: circSAMD4A targets and negatively regulates miR-29c-3p. (A) Upper panel, TargetScan analysis showing the putative recognition sequence of miR-29c-3p in circSAMD4A. Lower panel, a dual-luciferase reporter was used to verify the interaction between circSAMD4A and miR-29c-3p in 293 cells. The relative expression levels of (B) circSAMD4A and (C) miR-29c-3p were detected in SH-SY5Y cells transfected with sh-circ, pcDNA-circSAMD4A or matched controls via reverse transcription-quantitative PCR. **P<0.01, ***P<0.001. sh-circ, sh-circSAMD4A; miR, microRNA; circSAMD4A, circular RNA sterile α motif domain containing 4A; WT, wild-type; MUT, mutant; sh, short hairpin RNA; NC, negative control.
Article Snippet: miR-29c-3p mimics (including miR control, cat. no. HMI0439, concentration: ≥10 6 TU/ml), miR-29c-3p inhibitor (including miR-29c-3p inhibitor NC, cat. no. HSTUD0440, concentration: ≥10 6 TU/ml) were purchased from Sigma-Aldrich (Shanghai) Trading Co., Ltd., shNC (cat. no. C01001, concentration: 10 8 TU/ml), sh-circ (cat. no. C03002, concentration: 10 8 TU/ml),
Techniques: Sequencing, Luciferase, Expressing, Transfection, Real-time Polymerase Chain Reaction, Mutagenesis, shRNA, Negative Control

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A. Continuous sedimentation coefficient distribution for maize C4- and nonC4-NADP-ME at pH 8.0. The distribution profile for nonC4-NADP-ME shows variability in oligomeric states (dimers and tetramers) depending on the protein concentration. The protein concentration analyzed are indicated in the graphics. Data were analyzed using the c(s) model in the software package SEDFIT. B. Native PAGE (7%) analysis of 35 µg of protein extract from etiolated leaf and root tissue coupled to an in-gel NADP-ME activity assay. The excised protein bands and the corresponding identified NADP-ME isoforms are shown on the right. Recombinant C4- and nonC4-NADP-ME were used as controls (0.5 µg) and SERVA native marker is shown on the left. Bands with NADP-ME activity were analyzed by MS and the identified NADP-ME isoforms are listed on the right.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Sedimentation, Protein Concentration, Software, Clear Native PAGE, Activity Assay, Recombinant, Marker

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A. Overview of the chimeric and truncated N-terminal regions of the mature protein variants produced. B. Native PAGE (6%) of recombinant C4- and nonC4-NADP-ME variants with 5 μg protein loaded per lane followed by a Coomassie staining (left), or with 3 μg protein loaded per lane followed by an in-gel NADP-ME activity assay (right). C-D. Native PAGE (7%) of recombinant C4-NADP-ME ( C ) and Cyt2 ( D ) variants with 3 µg protein per lane followed by a Coomassie staining (left), or with 1 μg protein loaded per lane followed by an in-gel NADP-ME activity assay (right). In panels B-D, the SERVA native marker is shown on the left. E. Overview of the mutants produced at residue 200 in chimeric and truncated C4-NADP-ME variants and at the corresponding residue 208 in nonC4-NADP-ME.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Produced, Clear Native PAGE, Recombinant, Staining, Activity Assay, Marker, Residue

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A. Native PAGE (7%) analysis of recombinant nonC4- and C4-NADP-ME N-terminal variants, each with a point mutation at residues 148 or 140, respectively. Top panel: Immunoblot with 5 μg protein per lane using anti-His-HRP antibodies. Bottom panel: In-gel NADP-ME activity assay with 3 μg protein per lane. The SERVA native marker is shown on the right. B. Native PAGE (7%) analysis of recombinant nonC4-NADP-ME variants. Top panel: Coomassie staining with 3 μg protein per lane. Bottom panel: In-gel NADP-ME activity assay with 1 μg of protein per lane. Due to low protein recovery, 2.5 μg of IF and 1.5 μg Δ15 and Δ15IF were loaded. Recombinant C4- and nonC4-NADP-ME were used as controls. IF, AE, EQ, and FL correspond to the mutations I148F, A347E, E511Q, and F552L in nonC4-NADP-ME. In the nonC4-NADP-ME variant where 13 residues were simultaneously mutated (13aa), the enzyme contains the following mutations: F100T, I148F, H150N, D167N, R171T, N172D, E185V, R208G, Q209R, R274D, A347E, E511Q, and F552L. In the nonC4-NADP-ME variant where 20 residues were simultaneously mutated (20aa), the enzyme contains the following mutations: F100T, I148F, H150N, D167N, R171T, N172D, E185V, R208G, Q209R, R274D, H312D, I325F, A347E, V377M, H385Q, V482I, E511Q, N514T, D529A, and F552L (Suppl. Table 1).
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Clear Native PAGE, Recombinant, Mutagenesis, Western Blot, Activity Assay, Marker, Staining, Variant Assay

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A-B. Native PAGE (7%) analysis of recombinant C4-NADP-ME variants. Top panel: Coomassie staining with 3 μg protein per lane. Bottom panel: in-gel NADP-ME activity assay with 1 μg protein per lane. C. Continuous sedimentation coefficient distribution of C4G200R and C4D164N at pH 8.0. Data were analyzed using the c(s) model in the software package SEDFIT. D. Native PAGE (7%) analysis of recombinant nonC4-NADP-ME variants. Top panel: Coomassie staining with 3 μg of C4- and nonC4-NADP-ME and 7.5 µg of mutant proteins per lane. Bottom panel: in-gel NADP-ME activity assay with 1 μg of C4 and nonC4 controls, and 2.5 µg of mutant proteins per lane. The SERVA native marker is shown on the left.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Clear Native PAGE, Recombinant, Staining, Activity Assay, Sedimentation, Software, Mutagenesis, Marker

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A. Side view of the C4-NADP-ME structure (PDB ID: 5OU5), showing monomer A in pink and monomer B in purple. The C-D dimer is depicted in grey. Loops containing residues T163 and N164 are highlighted in yellow, while loops containing residue G200 in green. B. Top view of the dimer interface in C4-NADP-ME. Loops formed by residues T163 and D164 are highlighted in yellow. Loops containing residues 199-FGRPQG-204 in monomer A and 195-YGSIFGRPQG-204 in monomer B are shown in green. N-termini are indicated. C and D. Structural comparison of the loops at the dimer interface in C4-NADP-ME (C) and nonC4-NADP-ME (D) The differentially conserved substitutions in the loops are underlined for each isoform. E. Detailed view of the C4-NADP-ME dimer interface, emphasizing the loops in proximity to helix αA3 and the N-terminal helix αA1. For simplicity, only residues 96-101 of αA1 and 132-146 of αA3 in monomer A are shown. N- and C-termini of monomer A are indicated. The N-terminal β-sheet structures, which mediate interactions among all four monomers at the tetrameric interface, are enclosed within an oval. In B and E secondary structures of monomer A are shown in pink and those of monomer B in purple.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Residue, Comparison

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A. Cartoon illustration of the crystallographic structure of C4G200R. Monomer A (pink) and monomer B (purple) represent the dimer; likewise, monomer C (green) and monomer D (yellow). Pyruvate molecules at the four active sites and two dimeric interfaces are displayed as stick models in red. B. Top view of C4200R, displaying the tilted arrangement of the dimers relative to each other. C. 2D cryo-EM analysis of nonC4-NADP-ME and C4G200R. Representative 2D class averages of nonC4-NADP-ME at pH 8.0 (a), C4G200R at pH 8.0 (b) and C4-NADP-ME-G200R at pH 4.8 (c), respectively (Scale bar: 110 Å). The distribution (%) of oligomeric state (dimer; gray, tetramer; blue) of each sample is shown based on the number of single particles showing high resolution features for the respective oligomeric state.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Cryo-EM Sample Prep

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: A. Relative positioning of the N- and C-terminal ends, highlighting key loops containing residues T163 and D164 (yellow) and G200 (green) in monomers A and B of C4-NADP-ME. For simplicity, only monomers A (pink) and B (purple) are shown. The N- and C-termini, along with secondary structures near the tetramer interface, are indicated for monomer A. B. Polar interactions of D164 with R166, Y632, and K108 at the tetrameric interface of C4-NADP-ME. C. Polar interactions of N172 with L175, Y640, and N115 at the tetrameric interface of nonC4-NADP-ME. In panels B and C, black dashed lines indicate polar bonds between the interacting functional groups, with bond distances denoted in Å.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: Functional Assay

Journal: bioRxiv
Article Title: From dimer to tetramer: the evolutionary trajectory of C4 photosynthetic-NADP-ME oligomeric state in Poaceae
doi: 10.1101/2025.01.05.631420
Figure Lengend Snippet: The inset in the upper left corner shows the N-terminal sequences of maize Cyt2, nonC4-NADP-ME and C4-NADP-ME isoforms. Arrow sizes correspond to the relative prevalence of each oligomeric state, with question marks indicating the most probable state of the ancient isoforms. The scheme was designed as a free-form artistic illustration, with the lines not representing branch lengths.
Article Snippet: Expression vectors for various mutant variants of C4- and
Techniques: